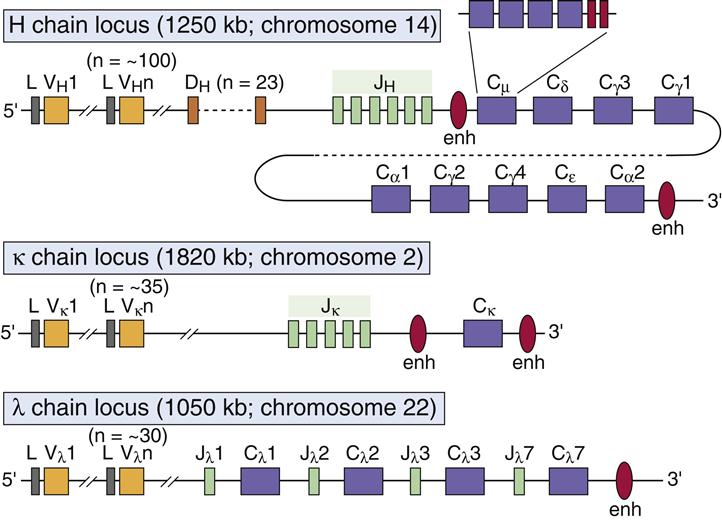
Mathematically, this strategy allows for ~6,000 possible heavy chain (subscript H) and ~300 possible light chain (subscript L) V(D)J combinations, for a total of ~1.8 million possible heavy-and-light chain pairings. The antigen-binding variable regions of antibody molecules draw combinatorially from a set of somatically encoded V, D, and J gene segments. Thus, as large-scale sequencing becomes cost-effective for clinical testing, we suggest that sequencing an individual's expressed antibody repertoire has the potential to become a useful diagnostic modality. We ran computer simulations to test whether enrichment for specific VDJ Hcombinations could be detected in "antigen-exposed" populations, and found that enrichment is detectable with moderate-to-high sensitivity and high specificity, even when some VDJ Hcombinations are not represented at all in some test sets. We find that for any two antigens chosen at random, chances are 90 percent that their repertoires' V Hsegments will overlap by less than half, and 98 percent that their VDJ Hcombinations will overlap by ≤10 percent. We find that repertoire overlap in V H-, D-, and J H-segment use is least for V Hsegments and greatest for J Hsegments, consistent with there being more V Hthan J Hsegments in the human genome. We do this by examining the overlap in V H-, D-, and J H- segment usage among sequenced repertoires. Specifically, we ask whether the large-scale sequencing of antibody repertoires might provide a diagnostic tool for detecting antigen exposure. Here we analyze the properties of all sequenced repertoires in order to better understand the specificity of antibody responses. 
To date several studies have sought to catalog the full suite of antibodies that humans naturally produce against single antigens or other specificities (repertoire).
0 Comments
Leave a Reply. |
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
